I am relatively new to Bokeh and I am currently trying to develop a data analytics dashboard with it. I was wondering if it were possible to deploy objects in a Bokeh dashboard which are created within R Studio. More specifically, I am trying to avail of the process mining package in R called bupaR, and use their animated process map function.
To provide an illustrative example of what I am trying to do, here is some code which builds a k-means clustering dashboard within Bokeh:-
import numpy as np
from sklearn import cluster, datasets
from sklearn.neighbors import kneighbors_graph
from sklearn.preprocessing import StandardScaler
from bokeh.io import curdoc
from bokeh.layouts import column, row
from bokeh.models import ColumnDataSource, Select, Slider
from bokeh.palettes import Spectral6
from bokeh.plotting import figure
np.random.seed(0)
# define some helper functions
def clustering(X, algorithm, n_clusters):
# normalize dataset for easier parameter selection
X = StandardScaler().fit_transform(X)
# estimate bandwidth for mean shift
bandwidth = cluster.estimate_bandwidth(X, quantile=0.3)
# connectivity matrix for structured Ward
connectivity = kneighbors_graph(X, n_neighbors=10, include_self=False)
# make connectivity symmetric
connectivity = 0.5 * (connectivity + connectivity.T)
# Generate the new colors:
if algorithm=='MiniBatchKMeans':
model = cluster.MiniBatchKMeans(n_clusters=n_clusters)
elif algorithm=='Birch':
model = cluster.Birch(n_clusters=n_clusters)
elif algorithm=='DBSCAN':
model = cluster.DBSCAN(eps=.2)
elif algorithm=='AffinityPropagation':
model = cluster.AffinityPropagation(damping=.9,
preference=-200)
elif algorithm=='MeanShift':
model = cluster.MeanShift(bandwidth=bandwidth,
bin_seeding=True)
elif algorithm=='SpectralClustering':
model = cluster.SpectralClustering(n_clusters=n_clusters,
eigen_solver='arpack',
affinity="nearest_neighbors")
elif algorithm=='Ward':
model = cluster.AgglomerativeClustering(n_clusters=n_clusters,
linkage='ward',
connectivity=connectivity)
elif algorithm=='AgglomerativeClustering':
model = cluster.AgglomerativeClustering(linkage="average",
affinity="cityblock",
n_clusters=n_clusters,
connectivity=connectivity)
elif algorithm=='KMeans':
model = cluster.KMeans(n_clusters= n_clusters)
model.fit(X)
if hasattr(model, 'labels_'):
y_pred = model.labels_.astype(int)
else:
y_pred = model.predict(X)
return X, y_pred
def get_dataset(dataset, n_samples):
if dataset == 'Noisy Circles':
return datasets.make_circles(n_samples=n_samples,
factor=0.5,
noise=0.05)
elif dataset == 'Noisy Moons':
return datasets.make_moons(n_samples=n_samples,
noise=0.05)
elif dataset == 'Blobs':
return datasets.make_blobs(n_samples=n_samples,
random_state=8)
elif dataset == "No Structure":
return np.random.rand(n_samples, 2), None
# set up initial data
n_samples = 1500
n_clusters = 2
algorithm = 'MiniBatchKMeans'
dataset = 'Noisy Circles'
X, y = get_dataset(dataset, n_samples)
X, y_pred = clustering(X, algorithm, n_clusters)
spectral = np.hstack([Spectral6] * 20)
colors = [spectral[i] for i in y]
# set up plot (styling in theme.yaml)
plot = figure(toolbar_location=None, title=algorithm)
source = ColumnDataSource(data=dict(x=X[:, 0], y=X[:, 1], colors=colors))
plot.circle('x', 'y', fill_color='colors', line_color=None, source=source)
# set up widgets
clustering_algorithms= [
'MiniBatchKMeans',
'AffinityPropagation',
'MeanShift',
'SpectralClustering',
'Ward',
'AgglomerativeClustering',
'DBSCAN',
'Birch',
'KMeans'
]
datasets_names = [
'Noisy Circles',
'Noisy Moons',
'Blobs',
'No Structure'
]
algorithm_select = Select(value='MiniBatchKMeans',
title='Select algorithm:',
width=200,
options=clustering_algorithms)
dataset_select = Select(value='Noisy Circles',
title='Select dataset:',
width=200,
options=datasets_names)
samples_slider = Slider(title="Number of samples",
value=1500.0,
start=1000.0,
end=3000.0,
step=100,
width=400)
clusters_slider = Slider(title="Number of clusters",
value=2.0,
start=2.0,
end=10.0,
step=1,
width=400)
# set up callbacks
def update_algorithm_or_clusters(attrname, old, new):
global X
algorithm = algorithm_select.value
n_clusters = int(clusters_slider.value)
X, y_pred = clustering(X, algorithm, n_clusters)
colors = [spectral[i] for i in y_pred]
source.data = dict(colors=colors, x=X[:, 0], y=X[:, 1])
plot.title.text = algorithm
def update_samples_or_dataset(attrname, old, new):
global X, y
dataset = dataset_select.value
algorithm = algorithm_select.value
n_clusters = int(clusters_slider.value)
n_samples = int(samples_slider.value)
X, y = get_dataset(dataset, n_samples)
X, y_pred = clustering(X, algorithm, n_clusters)
colors = [spectral[i] for i in y_pred]
source.data = dict(colors=colors, x=X[:, 0], y=X[:, 1])
algorithm_select.on_change('value', update_algorithm_or_clusters)
clusters_slider.on_change('value_throttled', update_algorithm_or_clusters)
dataset_select.on_change('value', update_samples_or_dataset)
samples_slider.on_change('value_throttled', update_samples_or_dataset)
# set up layout
selects = row(dataset_select, algorithm_select, width=420)
inputs = column(selects, samples_slider, clusters_slider)
# add to document
curdoc().add_root(row(inputs, plot))
curdoc().title = "Clustering"
And here is code which is executed in R Studio for an animated process map:-
library(bupaR)
library(processanimateR)
library(eventdataR)
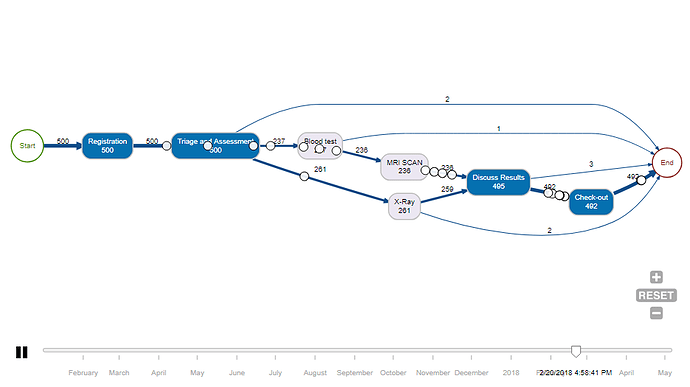
animate_process(patients)
Which give you this:-
Is there a way to deploy both of these features in a bokeh dashboard i.e. the process map to be beneath the clustering output or in a new tab? Or is there a more straightforward alternative to creating an animated process map within a bokeh dashboard that is more native to Python?
Many thanks in advance!