Hello, I am having a wired problem in the layout of the varea_stack plot:
as a source I have a pd DataFrame that looks a little like this:
budf0 budf1 ... budf9 budf10
0 2004-09-22 00:00:00 2004-09-22 00:00:00 ... 0.000000 8369.992188
1 2004-09-22 00:00:00 2004-09-22 00:00:00 ... 0.000000 8369.992188
2 2004-09-22 00:00:00 2004-09-22 00:00:00 ... -62263.421875 8369.992188
3 2004-09-22 00:00:00 2004-09-22 00:00:00 ... -60510.796875 8369.992188
4 2004-09-22 00:00:00 2004-09-22 00:00:00 ... -66217.226562 8369.992188
.. ... ... ... ... ...
792 2018-08-22 00:59:00 2018-08-22 00:59:00 ... 73861.562500 32149.527344
793 2018-09-01 00:25:00 2018-09-01 00:25:00 ... 73885.562500 32149.527344
794 2018-09-20 00:59:00 2018-09-20 00:59:00 ... 73959.929688 32149.527344
795 2018-09-21 00:59:00 2018-09-21 00:59:00 ... 73989.625000 32149.527344
796 2018-09-22 00:59:00 2018-09-22 00:59:00 ... 88514.390625 32149.527344
i want to do an varea_stack plot with “budf0” on the x axis and “budf3” - “budf10” as stackers, so my code looks like this:
stackPlot = figure()
stackPlot.xaxis.formatter = DatetimeTickFormatter()
stackNames = [f"budf{i}" for i in range(3,len(filteredDf.columns))]
stackPlot.varea_stack(stackers=stackNames, x="budf0", color=brewer['Spectral'][len(stackNames)], source=filteredDf)
show(stackPlot)
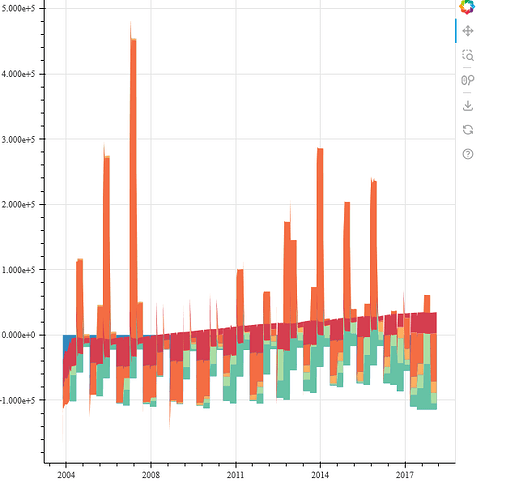
when i run this piece of code my plot looks like this:
the plot above is definitively not correct. (the graphs are just wrong)
However, when I plot each column of my source one after another like so:
budfs=["budf3","budf4","budf5","budf6","budf7","budf8","budf9","budf10"]
colo=brewer["Spectral"][11]
for i in range(len(budfs)):
y.varea_stack(stackers=[budfs[i]],x="budf0", color=colo[i], source=filteredDf)
show(y)
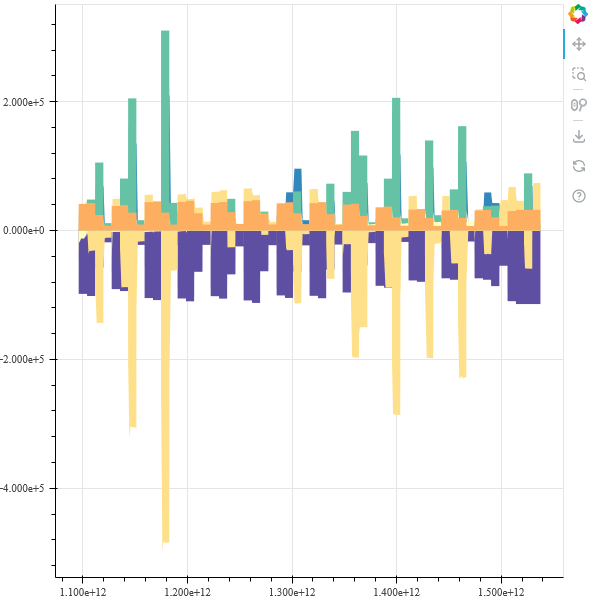
then the shape of the graph looks correct, like this:
But this is not varea_stack plot anymore, it is just each column plotted on top of each other. Can someone tell me what I do wrong? And why my original plot looks so wrong?
thanks very much in Advance